![]() Contributed by Anna Woodard, MD and Jeffrey A. Kant, MD, PhD
Contributed by Anna Woodard, MD and Jeffrey A. Kant, MD, PhD
CLINICAL PRESENTATION
A 62 year old woman presents to the clinic for neurologic and psychologic examination and genetic testing. Her cousin with mild chorea was recently diagnosed with Huntington disease (HD) following testing. Her maternal uncle is reported to have similar symptoms but has never undergone genetic testing. The patient's brother recently tested positive for HD as well at the age of 65. Her mother was diagnosed as a paranoid schizophrenic but never with this specific disorder. See Figure 1.
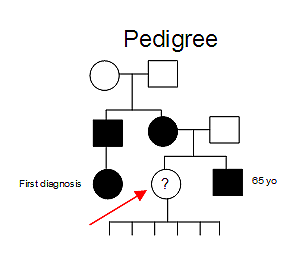
Figure 1; Patient's pedigree
The patient believes she has become more clumsy and has mild trouble with walking but denies any other symptoms. Her husband of several years does not notice any motor deficits. Physical exam and neurological exam are normal except for mild problems with tandem gait. The decision to proceed with genetic testing was reached following appropriate counseling and signed informed consent.
MOLECULAR TESTING
PCR Sizing for Assessment of Huntington Disease from UPMC Department of Pathology Division of Molecular Diagnostics [1].
Detection of an expanded trinucleotide CAG repeat in the first exon of the Huntingtin gene is based on PCR amplification of the Huntingtin gene using a pair of flanking primers. The 5' primer is fluorescently labeled and PCR fragment size is detected on an ABI capillary sequencer. An expanded CAG allele will produce a longer fragment than a normal allele. Four controls representing normal, intermediate, low and more highly expanded alleles are assayed along with the clinical samples. A negative control with no DNA is also assayed.
Successful DNA amplification and rough sizing is obtained by running the samples on an agarose gel stained subsequently with ethidium bromide. The gel should be examined to confirm the absence of amplification in the negative control. The PCR amplification products are then run on the ABI capillary sequencer and electropherograms are reviewed.
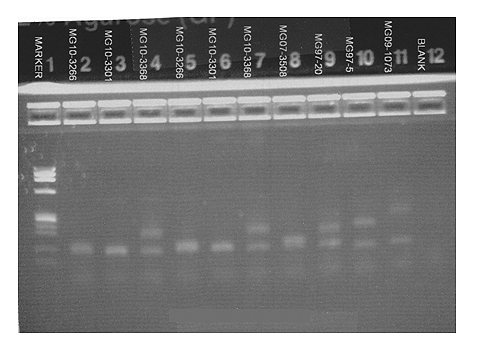
Figure 2: Gel electrophoresis of the control and patient samples. The patient sample is in lanes 4 and 7. Lane 9 contains the high normal control. Lanes 10 and 11 contain controls with one normal allele and one expanded allele.
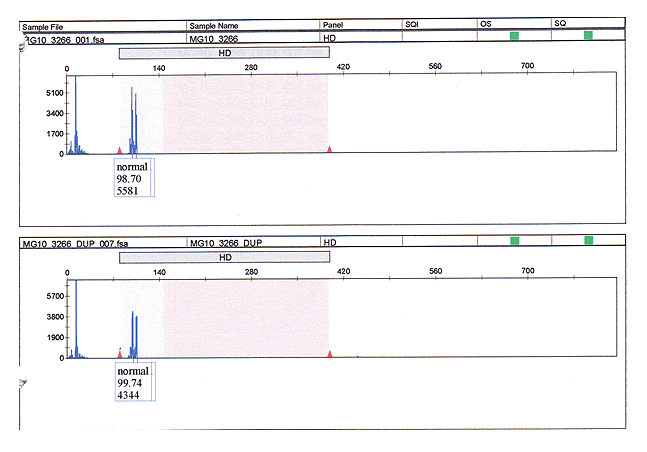
Figure 3: Normal control capillary sequencer results. This sample shows two peaks within the normal sequence range. The first peak represents 19 repeats and the second represents 21 repeats. The calculated CAG repeat lengths are the same for the replicate sample.
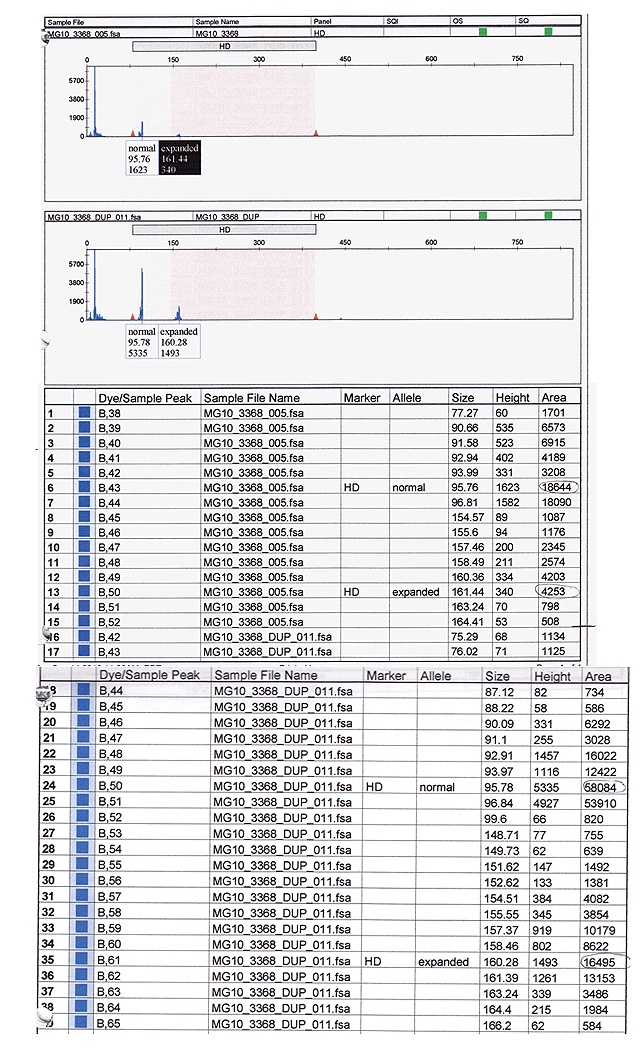
Figure 4: Patient capillary sequence results which show one peak within the normal range and one peak in the expanded range in both the upper and lower panels. Note that the area of the peaks is larger in the duplicate sample and these results were used to calculate the results in Table 1.
The number of CAG repeats is calculated using the following formula:
# CAG repeats= [(size of band in bp-48bp)/3]+2
This reflects subtraction of nucleotides which do not contribute to the CAG repeat region from the total length of PCR amplicons with the resulting value divided by 3 to give the number of CAG repeats. Values are rounded to the nearest whole number and have a "fudge factor" of +2 added to each value to compensate for undersizing associated with capillary electrophoretic analysis.
The calculated number of our patient's CAG repeats are reported in Table 1.

Table 1: The # of CAG repeats in the patient was calculated using the information from the lower panel of the capillary sequencing.
A normal size allele has a CAG repeat region of 9-26 repeats and an expanded allele is >35 repeats. An "intermediate" range of 27-35 repeats is also recognized.
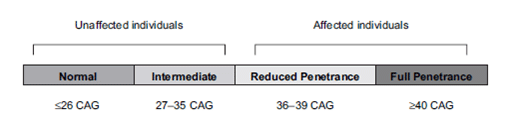
Figure 5: The current guidelines for penetrance dependent on #CAG repeats [2].