ASSAY PRINCIPLE AND INTERPRETATION:
Following genomic DNA isolation from peripheral blood leukocytes and its enzymatic digestion with PstI, fragments are separated using agarose gel electrophoresis, denatured, and transferred to a nylon membrane which is hybridized with a specific radioactively-labeled probe that recognizes a fragment containing the CGG repeat region upstream of the FMR1 gene. Analysis of the autoradiogram allows reasonably accurate sizing of CGG repeat expansions in the FMR1 gene based on the molecular weight of the fragment compared with marker DNA fragments that are run with patient samples. The Southern Blot is therefore the screening assay of choice to detect a wide range of FMR1 alles including rare individuals with full mutation expansions >200 repeats as well as those with premutation and even gray zone (45-55 repeats) alleles. It also allows detection of individuals with modest degrees of mosaicism.
Our patient's sample exhibits an expanded FMR1 allele size of ~2800 base pairs corresponding to ~570 CGG repeats with a minor species at ~2450 base pairs (~450 repeats). The presence of various sized alleles reflects the existence of different sized mutations in different populations of blood cells and is thought to result from varying degrees of CGG repeat number increase in different cells when the mother's premutation expands to a full mutation during embryogenesis. A normal sized allele is absent.
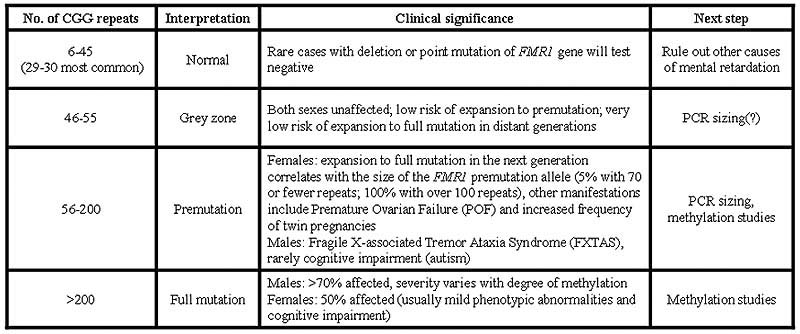
TABLE 1: CGG repeat numbers and their significance.

The definitive diagnosis of Fragile X syndrome is established via Southern Blot analysis using a methylation-sensitive restriction enzyme to probe methylation status of the region upstream of the FMR1 gene. Expansion of an unstable CGG repeat to over 200 triplets in the 5' untranslated region of the FMR1 gene leads to extensive local methylation of a CpG island just upstream from the repeat resulting in transcription silencing, loss of FMR1 protein and clinical features of Fragile X syndrome. The Southern blot analysis allows for the identification of the size of the (CGG)n repeat as well as the assessment of the methylation status of the CpG island. Two restriction enzymes are used for the digestion of genomic DNA: EcoR and methylation-sensitive Eag .
Our patient's sample is run with a healthy female control: the lower band represents the methylated inactivated X-chromosome, the upper band the unmethylated (active) X chromosome. Our patient has only one band (one X chromosome) that has a larger than normal molecular weight (due to presence of expanded CGG repeats) and is fully methylated, confirming the clinical diagnosis of Fragile X syndrome.