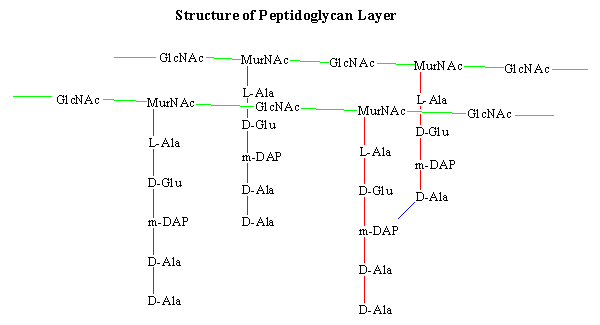
Figure 1: Peptidoglycan structure of gram-positive bacteria, showing D-Ala moiety crosslinking.
Vancomycin binds the D-Ala:D-Ala dipeptide, preventing the crosslinks from forming and
pentapeptide precursors from extending.
FINAL DIAGNOSIS: VANCOMYCIN-DEPENDENT ENTEROCOCCUS FAECIUM
DISCUSSION:
I) Background
The first documented strain of vancomycin-dependent enterococcus (VDE) was isolated in 1993 from the urine of a 46-year-old woman at Thomas Jefferson University Hospital in Philadelphia, Pennsylvania1. This patient had a prolonged and complicated hospitalization that required extended broad-spectrum antimicrobial therapy, including 123 days of vancomycin. Although her initial urine culture yielded vancomycin-resistant enterococci, subsequent urine specimens failed to yield organisms with standard culture techniques, despite gram-positive cocci in chains being seen on gram stain. Ultimately it was found that these gram-positive cocci had an extremely unusual nutritional requirement for growth - vancomycin. Subsequently, vancomycin-dependent E. faecalis (strain TJ310) was isolated from 5 additional urine cultures throughout the remainder of the hospital stay. This strain was eventually cleared with imipenem therapy.
Since this first isolate was reported, 20 additional cases of VDE have been reported worldwide, including both E. faecalis and E. faecium strains. Strains have been isolated from blood, urine, and stool and, in one case, a surgical wound infection. All VDE isolates followed vancomycin therapy (often prolonged), with the exception of one strain cultured from the blood, urine and dialysis catheter of a critically ill patient with no known prior exposure to vancomycin. Of the reported cases, VDE was considered pathogenic in only two. Also, vancomycin-dependent strains of enterococci have never been shown to affect the clinical course of any patient, including the patient described in this report. Although the clinical significance of VDE remains unclear, an outbreak of VDE in a bone marrow transplant unit at Johns Hopkins University in 1999 was reported2 and demonstrates its potential for becoming a clinically significant pathogen.
Numerous studies have shown, using pulsed-field gel electrophoresis that these strains exclusively evolve from vancomycin-resistant strains. Further molecular work on these bacteria showed that the acquisition of vancomycin resistance genes is necessary for mutation into the drug-dependent form. In fact, VDE might best be considered a mutant VRE. Although this patient did not grow VRE on prior rectal swabs, the acquisition of resistance genes may have occurred in the 7 days separating her VDE culture from her most recent negative rectal swab. This phenomenon of progressing from susceptibility to tolerance to dependence is best understood at the molecular genetic level of both vancomycin sensitive and resistant enterococcal species.
II) Vancomycin resistance mechanism
Figure 1 illustrates the peptidoglycan structure in Gram-positive bacteria. The D-Ala:D-Ala subunits of the pentapeptide precursors form crosslinks with moieties on neighboring pentapeptides, ultimately giving the peptidoglycan layer its structure. In bacteria, the ddl gene codes for the ligase enzyme responsible for terminal D-Ala:D-Ala formation.

Vancomycin binds the terminal D-Ala:D-Ala structure, preventing crosslinks from forming and pentapeptide structures from extending during synthesis. Resistance to vancomycin is mediated by a plasmid gene cluster that codes for seven proteins, 6 of which have been extensively studied and are well understood4. Two distinct resistance phenotypes have been identified: VanA - resistant to high levels of vancomycin (MIC, 64 to > 1000 ug/ml) and teicoplanin (MIC, 16 to 512 ug/ml), and VanB - resistant to various levels of vancomycin (MIC, 4 to > 1000 ug/ml) and susceptible to teicoplanin. Both of these resistance phenotypes have been identified in vancomycin dependent strains. The genes responsible for resistance in both phenotypes, as well as their associated proteins are listed below in Table 1.
This gene cluster, in both phenotypes, acts to a) detect the presence of vancomycin and start transcription of specific resistance genes, b) form and incorporate D-Ala:D-Lac into the growing peptidoglycan wall, and c) eliminate any D-Ala:D-Ala precursors, thereby eliminating the vancomycin sensitive pathway of peptidoglycan formation. The D-Ala:D-Lac dipeptide is fully capable of forming cross-links with neighboring pentapeptides; however vancomycin cannot bind this dipeptide structure and thus has no effect cell wall synthesis. Important to both vancomycin resistant and dependant strains of enterococcus is the fact that the resistance gene cluster is not constitutively expressed, but induced by the presence of antibiotic.
III) Vancomycin dependence - proposed mechanism
Molecular studies performed on both the original isolate, TJ310, as well as subsequent isolates have shown that primary defect conferring vancomycin dependence in these strains involves mutations in the ddl gene, which encodes the enterococcal D-Ala:D-Ala ligase protein5,6. Mutations in the ddl gene in the VDE strains were shown to result in the synthesis of dysfunctional, truncated proteins or proteins with amino acid substitutions that resulted in inadequate substrate binding. With the absence of functional ligase activity, these isolates require glycopeptide induction of the resistance pathway for cell wall synthesis.
Gene sequencing of several VDE strains, as well as a VRE strain (TJ282) isolated from the same patient as TJ310 and a control VRE strain (V583) were performed. The ddl sequences from strains TJ282 and E. faecalis V583 were identical. The VDE isolates showed point mutations leading to amino acid substitutions, nucleotide repeat insertions causing frameshift mutations, and deletions resulting in loss of more than one amino acid. More specifically, each strain had either a 5-bp insertion at codon 41 resulting in an early stop codon, an in-frame 6-bp deletion resulting in the loss of two residues (KDVA243-246->KA), or single amino acid substitutions (E13->G, G99->R, V241->D, D295->G, P313->L)4. Many of the mutations that resulted in amino acid substitution or deletion occurred in regions of the enzyme known to be critically involved in substrate binding. Subsequent biochemical analysis confirmed that VDE did in fact have impaired D-Ala:D-Ala ligase activity. In addition to these mutations, other mutations in the dll gene have also been identified.
IV) Revertant Mutants
The inability of VDE to survive in the absence of vancomycin makes them seemingly easy to eliminate - discontinue the drug. Surprisingly, withdrawal of vancomycin (in in-vitro studies) does not always eliminate the enterococcus strain but rather results in the unusual phenomenon of revertant mutation5. In this phenomenon, these bacteria become senescent and undergo additional genetic change that actually restores the D-Ala:D-Ala ligase function, resulting in another genetically distinct vancomycin-resistant strain. In essence what develops is cyclic mutational change that causes the bacteria to shift between resistance and dependance phenotypes.
A multi-institutional study in 1999 elucidated the genetic mechanism by which revertant mutation occurs. The ddl gene was sequenced in several strains of VDE (including TJ310) and their VRE revertant mutants. This study uncovered two interesting facts. In vancomycin-resistant, teicoplanin-susceptible revertants, there had actually been base pair insertions in the deranged ligase gene that resulted in amino acid insertions in positions 251-253. One of the amino acids, Lys 253, is thought to be essential for enzyme activity. In the vancomycin and teicoplanin- resistant revertants, no sequence alterations were detected, indicating that the resistance genes may have been constitutively expressed.
Although these studies indicate that withdrawal of vancomycin may not be adequate to eliminate these strains, revertant mutation has not been detected in clinical situations.
V) Summary
Vancomycin-dependent enterococci illustrate the power of biologic adaptation. Not only do these bacteria acquire the ability to actually utilize vancomycin for cell wall synthesis, but they also have the ability to revert to their resistant phenotypes when the drug is withdrawn. Interestingly, thus far these organisms are more of a curiosity than a significant clinical pathogen. Nevertheless, the outbreak of VDE at Johns Hopkins University in 1999 suggests that these bacteria do have potential for creating another layer of iatrogenic infectious disease complexity. At the very least, this organism beautifully exemplifies the perpetual battle between clinicians and micro organisms.
REFERENCES:
![]() Contributed by Ken Clark, MD and A William Pasculle, ScD.
Contributed by Ken Clark, MD and A William Pasculle, ScD.